Reports of research work funded by grants prior to 2015
Malaghan Institute of Medical Research
Harnessing new technologies for optimal vaccine-induced antibody responses
H Poyntz
Gut Inflammation Research Group
Project Objective
We have generated two lines of BALB/c mice that mount divergent antibody production in response to vaccination. Using microbial transfer techniques we have identified gut microbiota to be the critical factor regulating the magnitude of these antibody responses. Utilising these unique strains of mice, we aimed to identify the pathways that dictate optimal antibody response through identification of the immune mechanisms that are manipulated by the gut microbiota.
The grant funded a panel of fluorescently tagged antibodies that could be used with a confocal microscope to image fixed or live cells isolated from draining lymph nodes of vaccinated mice, enabling us to see which cells are interacting to form the antibody response. Once we had identified the cell types of interest and the optimal time point, we planned to sort these cells from the lymph nodes and quantify expression levels of genes involved in provision of T cell help to B cells, differentiation, antibody production and antibody class switching within the B cells. For this, the grant funded the primers that are targeted to our genes of interest and these were used with quantitative polymerase chain reaction (qPCR) to determine expression levels.
Methods
Optimising a multi-colour panel for confocal microscopy:
Mice from two lines of BALB/c mice were vaccinated and lymph nodes were taken 14 days later and frozen in OCT then sectioned. Fluorochrome-conjugated antibodies were used to identify cell lineage and activity: CD4, B220, GL7, AID, CD40L.
Identifying critical pathways in the induction of an optimal antibody response:
Mice from two lines of BALB/c mice were vaccinated and specific T cell lineages sorted from lymph nodes seven days later. RNA was isolated and converted to cDNA before qPCR determination of gene expression. For whole lymph node gene expression analysis, RNA was isolated from draining lymph nodes 14 days after vaccination.
Results
We have determined that at day 14 after vaccination the germinal centre reaction, key to driving the antibody response, had formed in both lines of BALB/c mice. A panel of three antibodies to visualise germinal centre structure, locality and quantify their number was successfully determined through multiple rounds of optimisation (Fig. 1A). The quantity of germinal centres throughout the lymph node structure was determined using this staining panel (Fig. 1B).
Day seven after vaccination was determined to be the peak of the CD4+ T cell response to vaccination. Follicular T helper cells (TFH) and extra follicular T helper cells (EFH) were sorted from lymph nodes and probed for the expression of genes critical in the provision of T cell help to B cells; IL-4, IL-21 and Bcl6 (Fig 2 A, B, C). The expression level of genes involved in antibody production and antibody class switching within B cells was measured 14 days after vaccination; AID, BLIMP1 (Fig 2 D, E), April, BAFF, IL-6, Bcl6 (data not shown).
Conclusions
We hypothesised that line A, the low antibody responders may have a reduced number of germinal centres or a disrupted germinal centre structure, which would impede optimal generation of antibody producing plasma cells. Preliminary results indicate no difference in germinal centre structure between the two lines of mice but there is a trend to a reduced number of germinal centres in line A. This method of quantification is continuing.
Antibodies of IgG1 and IgG2a isotype are induced by our vaccine regime. TFH provide help to B cells for antibody production in response to vaccination. That Bcl6, the transcription factor required for TFH differentiation, is up regulated in TFH of line B indicates there may be greater differentiation of CD4+ T cells to the TFH subset in these mice. Greater expression levels of IL-4 and IL-21 in TFH of line B suggests more stimulation is provided to the B cells in line B compared to line A. Activation induced cytokine deaminase (AID) is expressed in B cells to mediate class-switching of the antibody isotype produced by the B cell. BLIMP1 is a transcription factor that promotes differentiation of B cells to a plasma cell phenotype. That expression of both these factors is increased in lymph nodes of line B indicates class-switching and plasma cell differentiation may be enhanced in line B as compared to line A. The next step will be to sort germinal centre B cells and plasma cells from the lymph nodes for a targeted analysis of gene expression within these cells.
This grant has allowed the optimisation of methods implementing new technologies and provided data to delineate key immunological mechanisms that are differentially regulated in these two lines of mice. The project will continue to identify the signalling pathways required to drive optimal antibody responses and understand how the microbiota is manipulating these pathways to influence optimal antibody responses.
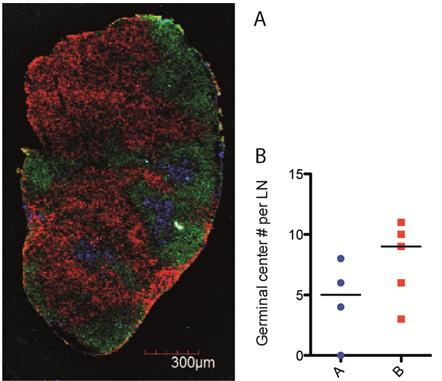
Fig. 1 Germinal centre formation in lymph nodes of two lines of BALB/c mice. A Image taken on confocal microscope at 4000x magnification. Red CD4 (CD4+ T cells), Green B220 (B cells) and blue GL7 (Germinal centre). B Quantification of germinal centre number in lymph node sections. 5 mice per group.
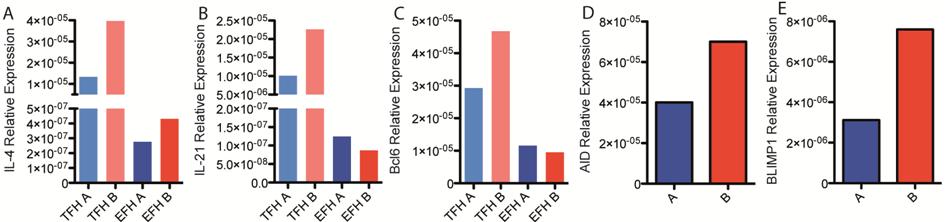
Fig. 2 A – C Expression of genes involved in provision of T cell help to B cells. Gene expression in sorted populations of TFH and EFH relative to 18S control. Sample pooled from 6 mice per group. D – E Expression of genes involved in antibody production within B cells. Gene expression relative to 18S control in total cells of draining lymph node. Mean of 4 mice per group displayed.



