Reports of research work funded by grants prior to 2012
Victoria University of Wellington
Developing Enzymes to Image and Target Tumours by Gene Therapy
JN Copp and DF Ackerley
School of Biological Sciences
Project Overview
Tumour-targeting viruses and bacteria hold great promise as anti-cancer agents. They kill cells by entirely different mechanisms to radio- and chemotherapies, and have potential to synergise with these treatments without overlapping toxicities. Furthermore, these agents can be ‘armed’ with genes that encode enzymes that activate prodrugs – compounds that are inert in their administered form, but become highly toxic upon activation. This not only improves killing of infected cells, but also neighbouring non-infected cells, as the activated prodrug can diffuse locally and exert a ‘bystander effect’. A critical limitation to date has been the inability to non-invasively monitor the location and amplification of viral and bacterial vectors in the body post-administration. To address this, we are using directed evolution (random genetic mutagenesis coupled with artificial selection for improved enzyme variants) to develop optimised Positron Emission Tomography (PET) imaging capable nitroreductases that also have enhanced prodrug activation and biosafety potential. These enzymes will ultimately facilitate uptake of enzyme-prodrug gene therapy into clinical trials, and accelerate development of self-replicating oncolytic biological agents.
Progress to Date
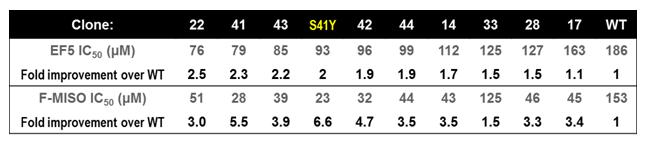
Specific Objective 1: To optimise the ability of previously developed prodrug-activating enzymes to co-metabolise probes with PET imaging capabilities. We previously used directed evolution to engineer variants of the nitroreductase enzyme NfsA from Escherichia coli with improved ability to activate the dinitro-benzamide prodrug PR-104A. Here, these variants were tested for their ability to co-metabolise two preferred PET imaging probes, etanidazole pentafluoride (EF5) and fluoromisonidazole (F-MISO). Serendipitously it was discovered that a single residue mutation from wild type NfsA that had a dominant effect on improved PR-104A metabolism (S41Y) also had a substantially dominant effect on the ability to activate the 2-nitroimidazole PET probes EF5 and F-MISO (Table 1). Thus, all NfsA variants previously selected for improved PR-104A metabolism were also substantially improved at activating preferred PET probes. We hypothesise that this will provide sufficient sensitivity that human tumours infected with viral or bacterial vectors “armed” with genes encoding these evolved nitroreductases will not only be more sensitive to PR-104A prodrug, but also visible in PET scans using EF5 or F-MISO as probe. This hypothesis will be tested in a future project via collaboration with researchers at the MAASTRO radiation oncology clinic in Maastricht, the Netherlands.
Table 1: Improved metabolism of 2-NI PET capable probes by evolved nitroreductases.
E. coli cells expressing evolved nitroreductase variants were sensitive to lower doses of 2-NI probes than cells expressing the wild type NfsA enzyme (WT), due to the dominant effect conferred by the S41Y single-residue mutation that was present in all clones apart from WT.
Specific Objective 2: To optimise a nitroreductase enzyme specifically for maximum PET imaging. This aspect of the study has sought to evolve nitroreductase enzyme variants specifically for optimal PET imaging capabilities, independent of the ability to activate prodrugs. An optimised F-MISO or EF5 activating nitroreductase would have considerable value as a PET reporter gene to promote development of next-generation GDEPT vectors and independent oncolytic agents. To achieve this, we have carried out random and targeted mutagenesis of E. coli NfsA to recover single-residue mutants with enhanced ability to activate RB6145, a genotoxic 2-NI analogue of EF5 and F-MISO (based on our genetic screening technology being responsive to DNA damage induced by activated RB6145, but not activated EF5 or F-MISO directly). We have recovered mutations at 12 residue locations that have yielded improvement in the ability to activate EF5 and/or F-MISO. These mutations are now being combined and re-screened to recover enzyme variants bearing synergistic combinations of the individual point mutations. To date indications are that we will succeed in generating nitroreductase variants that exhibit far greater improvement in 2-NI PET probe metabolism (relative to wild type NfsA) than any of the prodrug-optimised variants listed in Table 1.
Specific Objective 3: To optimise a nitroreductase enzyme for the ability to co-metabolise a promising next-generation prodrug, a PET imaging compound, and a bio-safety molecule that has the ability to suppress replication of bacterial and viral vectors. Research toward this objective is ongoing.



